Summarizing Embeddings for Administrative Regions
Source:vignettes/region-summaries.Rmd
region-summaries.RmdIntroduction
When working with large geographic regions, you often need summary
statistics rather than full pixel-level embeddings. The
summarize_region() method provides a pipeline that:
- Downloads embeddings to a temporary location
- Optionally masks pixels to match irregular region boundaries
- Computes user-specified summary statistics
- Automatically cleans up downloaded data
This vignette demonstrates how to use this functionality with administrative boundaries from a shapefile.
Demonstration with Simulated Data
Before working with real data, let’s demonstrate the summarization workflow with simulated embeddings and regions.
Creating Example Regions
# Create 4 example polygon regions (simulating LGAs)
region1 <- st_polygon(list(matrix(c(
0, 0, 1, 1, 0,
0, 1, 1, 0, 0
), ncol = 2))) |> st_sfc(crs = 4326)
region2 <- st_polygon(list(matrix(c(
1, 1, 2, 2, 1,
0, 1, 1, 0, 0
), ncol = 2))) |> st_sfc(crs = 4326)
region3 <- st_polygon(list(matrix(c(
0, 0, 1, 1, 0,
1, 2, 2, 1, 1
), ncol = 2))) |> st_sfc(crs = 4326)
region4 <- st_polygon(list(matrix(c(
1, 1, 2, 2, 1,
1, 2, 2, 1, 1
), ncol = 2))) |> st_sfc(crs = 4326)
# Combine into sf object
regions <- st_sf(
name = c("Region A", "Region B", "Region C", "Region D"),
land_type = c("urban", "agricultural", "forest", "mixed"),
geometry = c(region1, region2, region3, region4)
)
# Plot the regions
ggplot(regions) +
geom_sf(aes(fill = land_type), alpha = 0.7) +
geom_sf_text(aes(label = name), size = 4) +
scale_fill_manual(values = c("urban" = "#E41A1C",
"agricultural" = "#FFD92F",
"forest" = "#4DAF4A",
"mixed" = "#984EA3")) +
labs(title = "Example Administrative Regions",
subtitle = "Simulating 4 LGAs with different land cover types") +
theme_minimal()
#> Warning in st_point_on_surface.sfc(sf::st_zm(x)): st_point_on_surface may not
#> give correct results for longitude/latitude data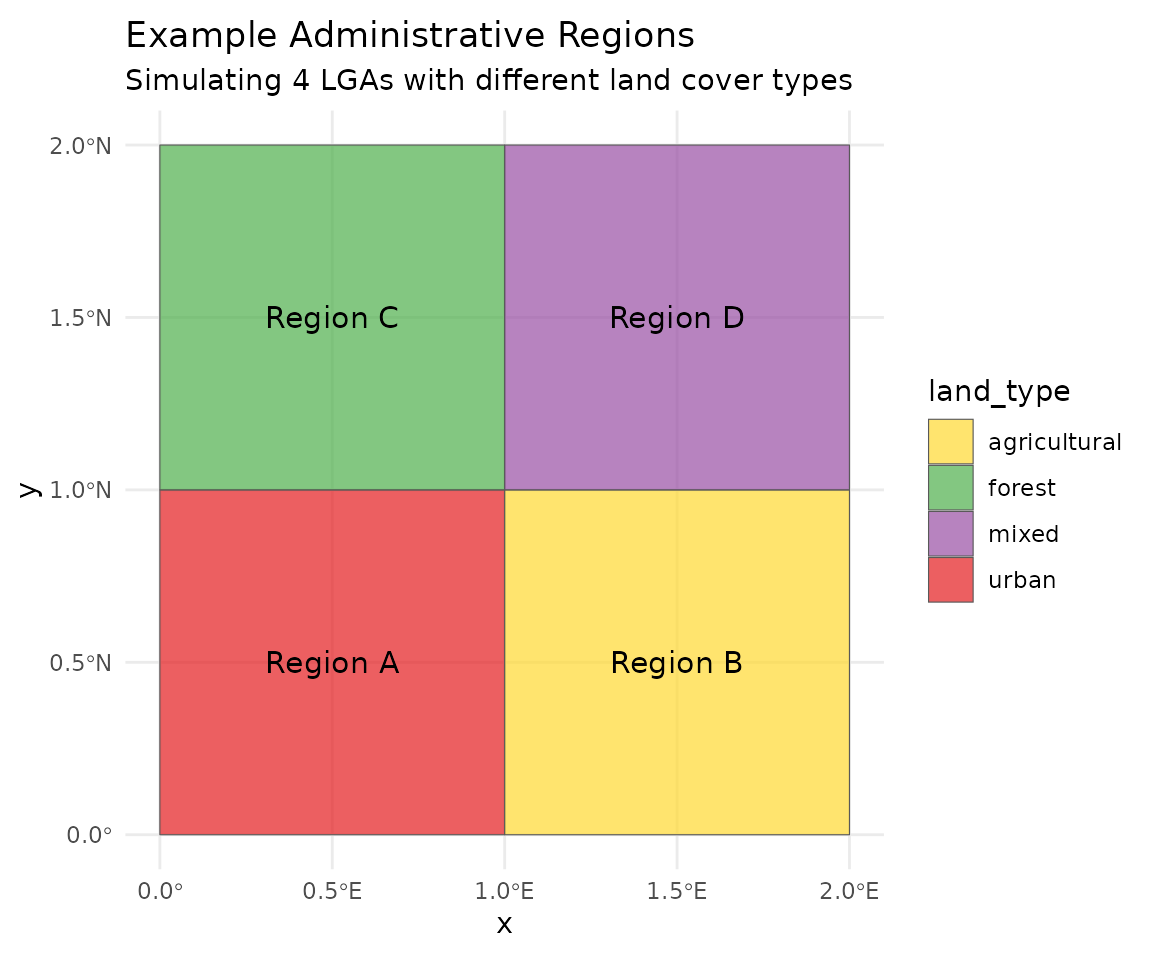
Simulating Region Embeddings
set.seed(42)
# Simulate mean embeddings for each region type
# Different land types have distinct embedding signatures
simulate_embedding <- function(land_type) {
base <- switch(land_type,
"urban" = rnorm(128, mean = 0.5, sd = 0.1),
"agricultural" = rnorm(128, mean = 0.0, sd = 0.15),
"forest" = rnorm(128, mean = -0.4, sd = 0.12),
"mixed" = rnorm(128, mean = 0.1, sd = 0.2)
)
base
}
# Generate embeddings for each region
region_embeddings <- lapply(regions$land_type, simulate_embedding)
names(region_embeddings) <- regions$name
# Create embedding matrix
emb_matrix <- do.call(rbind, region_embeddings)
rownames(emb_matrix) <- regions$name
cat("Embedding matrix dimensions:", dim(emb_matrix), "\n")
#> Embedding matrix dimensions: 4 128
cat("Rows:", rownames(emb_matrix), "\n")
#> Rows: Region A Region B Region C Region DRegion Similarity Heatmap
# Calculate cosine similarity between regions
cosine_sim <- function(a, b) {
sum(a * b) / (sqrt(sum(a^2)) * sqrt(sum(b^2)))
}
n_regions <- nrow(emb_matrix)
sim_matrix <- matrix(0, n_regions, n_regions)
for (i in 1:n_regions) {
for (j in 1:n_regions) {
sim_matrix[i, j] <- cosine_sim(emb_matrix[i, ], emb_matrix[j, ])
}
}
rownames(sim_matrix) <- colnames(sim_matrix) <- regions$name
# Convert to data frame for plotting
sim_df <- as.data.frame(as.table(sim_matrix))
names(sim_df) <- c("Region1", "Region2", "Similarity")
ggplot(sim_df, aes(x = Region1, y = Region2, fill = Similarity)) +
geom_tile() +
geom_text(aes(label = round(Similarity, 2)), color = "white", size = 4) +
scale_fill_gradient2(low = "blue", mid = "white", high = "red", midpoint = 0.5) +
labs(title = "Region Similarity Matrix",
subtitle = "Cosine similarity based on mean embeddings") +
theme_minimal() +
theme(axis.text.x = element_text(angle = 45, hjust = 1))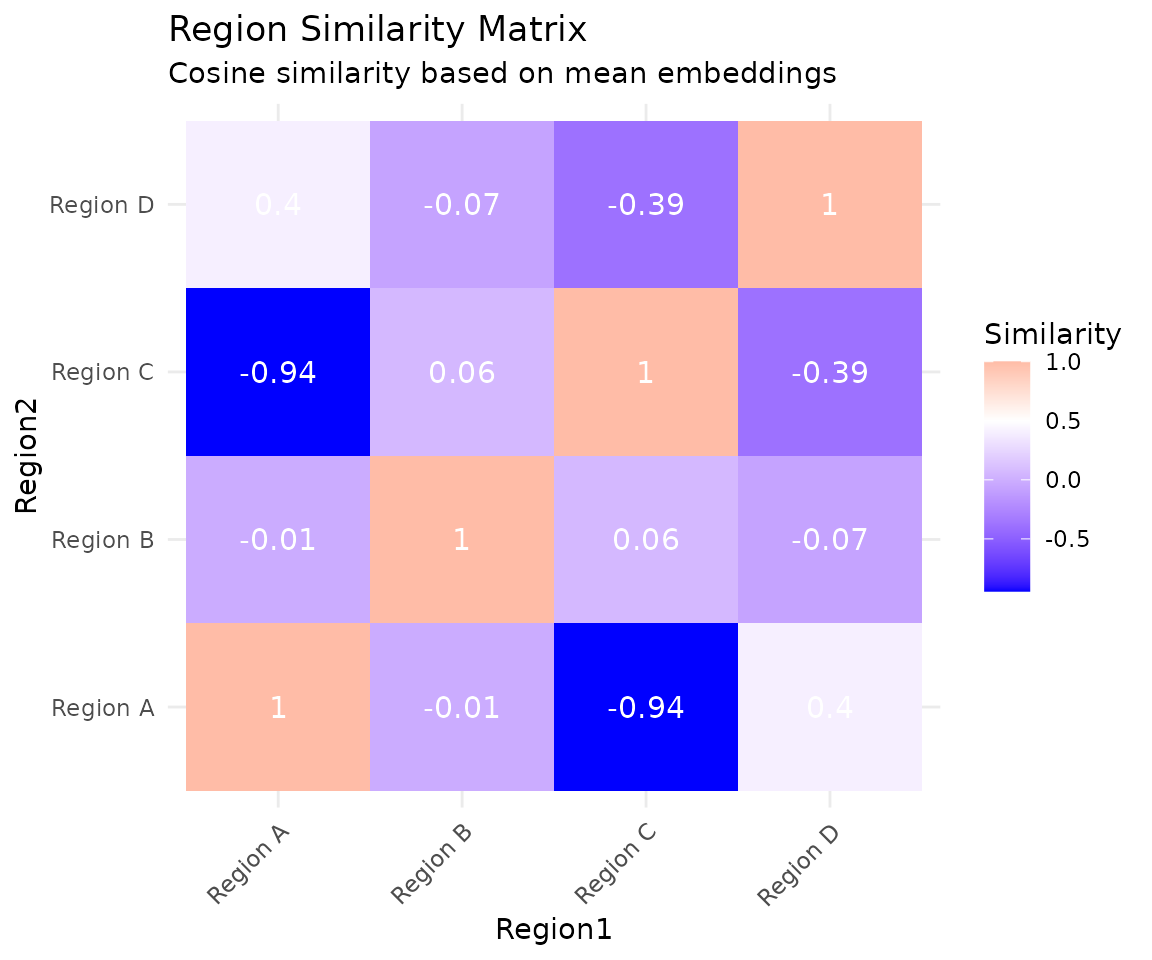
Embedding Profile Comparison
# Compare first 20 dimensions across regions
profile_df <- data.frame(
Dimension = rep(1:20, 4),
Value = c(emb_matrix[1, 1:20], emb_matrix[2, 1:20],
emb_matrix[3, 1:20], emb_matrix[4, 1:20]),
Region = rep(regions$name, each = 20)
)
ggplot(profile_df, aes(x = Dimension, y = Value, color = Region)) +
geom_line(linewidth = 1) +
geom_point(size = 2) +
scale_color_brewer(palette = "Set1") +
labs(title = "Embedding Profiles by Region",
subtitle = "First 20 of 128 dimensions shown",
x = "Embedding Dimension", y = "Value") +
theme_minimal() +
theme(legend.position = "bottom")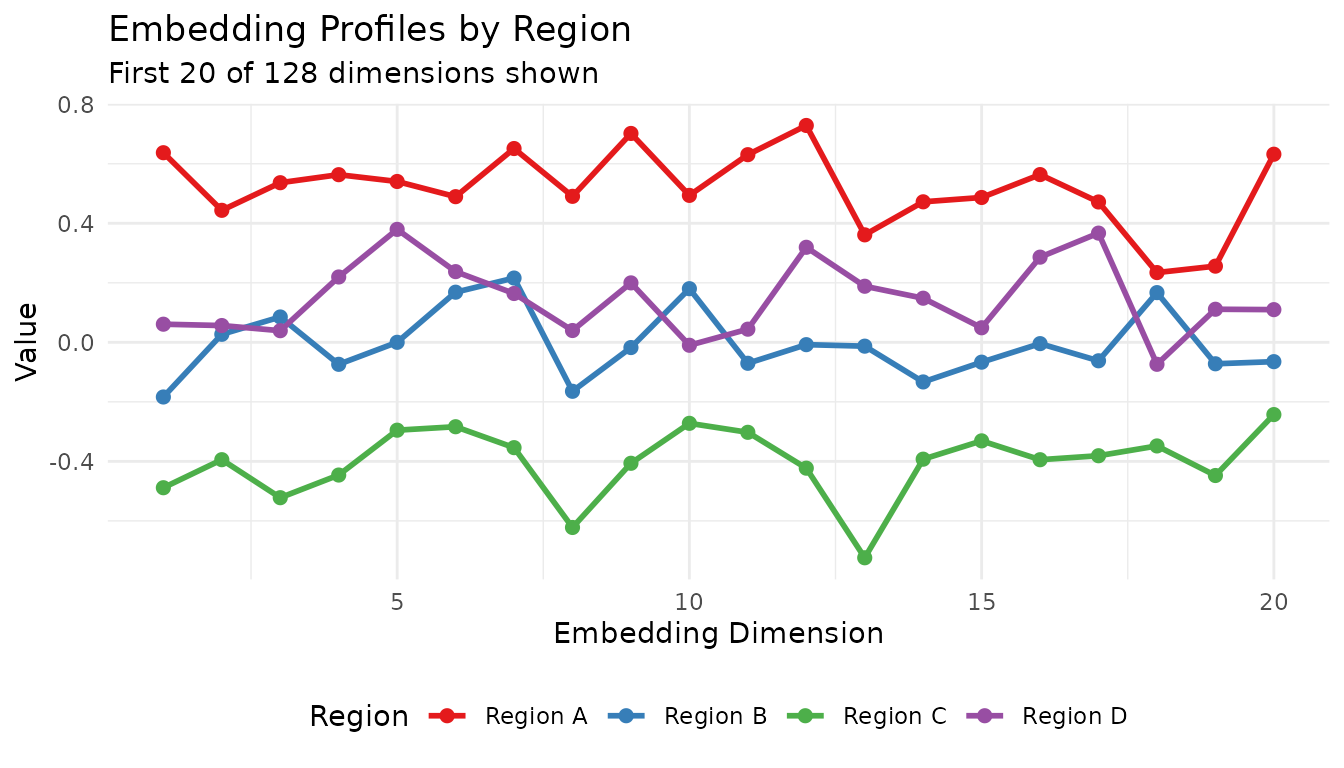
Hierarchical Clustering of Regions
# Cluster regions by embedding similarity
dist_matrix <- dist(emb_matrix)
hc <- hclust(dist_matrix, method = "ward.D2")
# Simple dendrogram plot
plot(hc, main = "Hierarchical Clustering of Regions",
sub = "Based on 128-dimensional embeddings",
xlab = "", ylab = "Distance")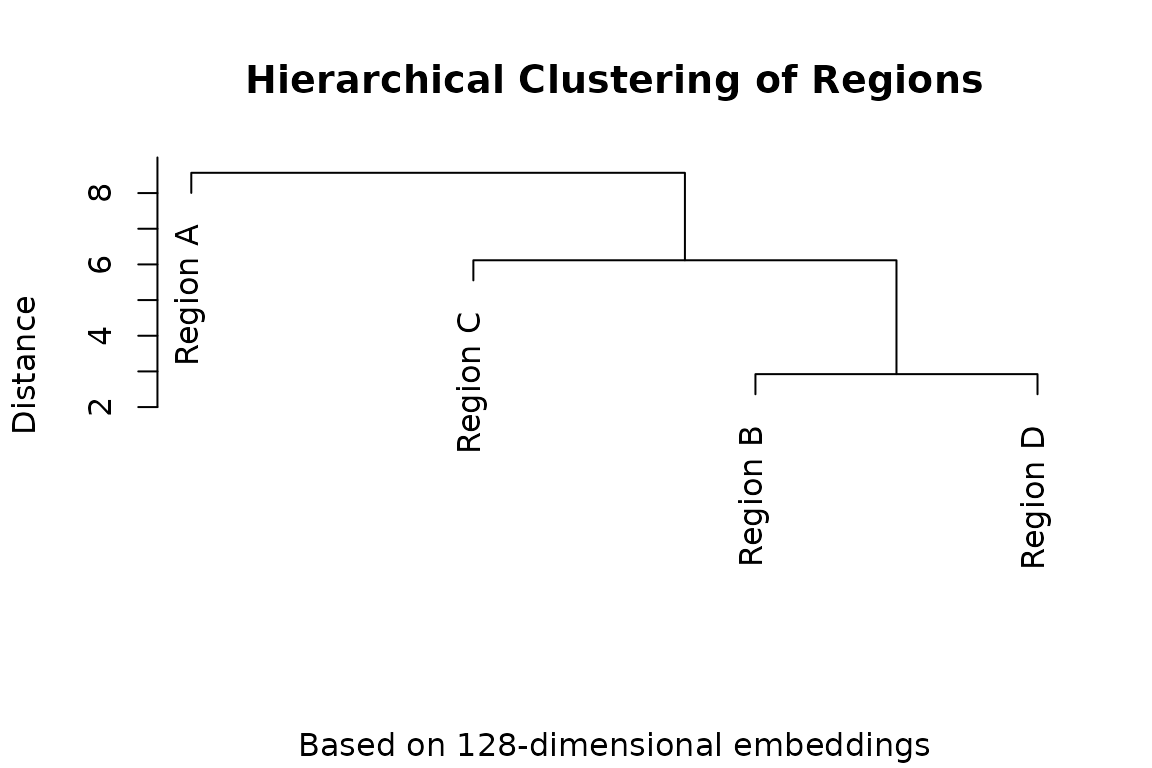
Map with Cluster Assignment
# Cut tree into 2 clusters
regions$cluster <- factor(cutree(hc, k = 2))
ggplot(regions) +
geom_sf(aes(fill = cluster), alpha = 0.7) +
geom_sf_text(aes(label = paste0(name, "\n(", land_type, ")")), size = 3) +
scale_fill_brewer(palette = "Set2") +
labs(title = "Regions Clustered by Embedding Similarity",
subtitle = "Regions with similar land cover cluster together") +
theme_minimal()
#> Warning in st_point_on_surface.sfc(sf::st_zm(x)): st_point_on_surface may not
#> give correct results for longitude/latitude data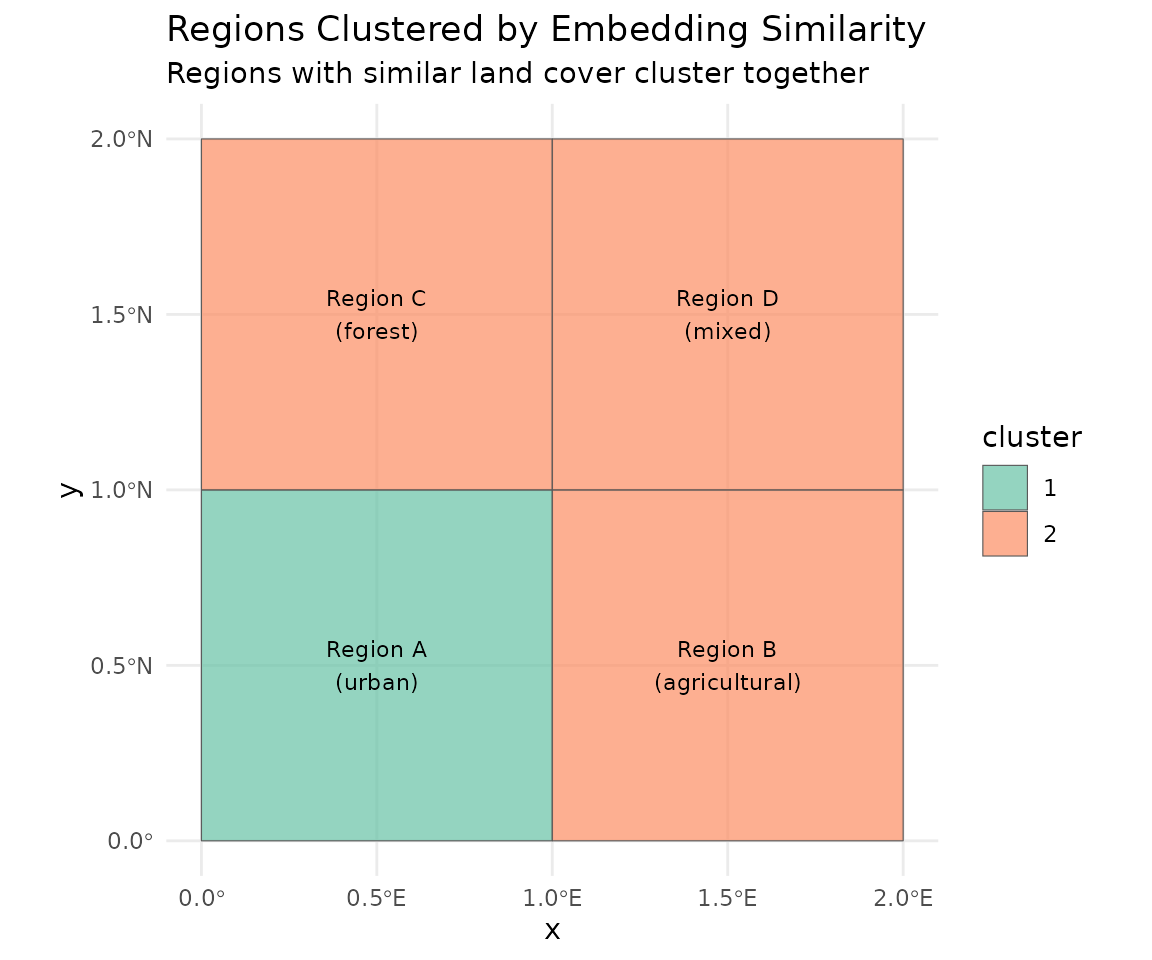
Loading Administrative Boundaries
We’ll use Nigeria’s Local Government Areas (LGAs) as an example. This shapefile contains 774 administrative units at the admin2 level.
# Load the shapefile
nigeria_lgas <- st_read("shapefiles/nigeria_admin2_shapefile.shp", quiet = TRUE)
# Examine the structure
print(nigeria_lgas[1:5, c("adminCode", "adminName")])Simple feature collection with 5 features and 2 fields
adminCode adminName geometry
1 NG001001 Aba North MULTIPOLYGON (((7.401109 5....
2 NG001002 Aba South MULTIPOLYGON (((7.387495 5....
3 NG015001 Abaji MULTIPOLYGON (((7.045872 9....
4 NG003001 Abak MULTIPOLYGON (((7.811244 5....
5 NG011001 Abakaliki MULTIPOLYGON (((8.4109 6.28...Summarizing a Single LGA
Let’s compute embedding summaries for Aba North, a small LGA in southeastern Nigeria.
# Select Aba North LGA
aba_north <- nigeria_lgas[nigeria_lgas$adminName == "Aba North", ]
# View the region
plot(st_geometry(aba_north), main = "Aba North LGA", col = "lightblue")Basic Summary
The summarize_region() method accepts an sf object
directly. It will:
- Use the bounding box to determine which tiles to download
- Mask pixels outside the polygon boundary
- Compute summary statistics
# Create GeoTessera client
gt <- geotessera()
# Compute summaries for Aba North
result <- gt$summarize_region(
region = aba_north,
year = 2024
)
# View the structure
names(result)
# [1] "summaries" "metadata"
# View metadata
result$metadata
# $region_bbox
# [1] 7.331234 5.081948 7.402708 5.134542
#
# $year
# [1] 2024
#
# $n_tiles
# [1] 4
#
# $is_masked
# [1] TRUEDefault Summary Statistics
By default, summarize_region() computes three
summaries:
# Mean embedding (128-dimensional vector)
mean_embedding <- result$summaries$mean
length(mean_embedding)
# [1] 128
# First 10 dimensions
head(mean_embedding, 10)
# Centroid embedding (sampled at geographic center)
centroid_embedding <- result$summaries$centroid
length(centroid_embedding)
# [1] 128
# Coverage statistics
result$summaries$coverage
# n_tiles n_valid_pixels n_total_pixels coverage_percent
# 4 xxxxx xxxxx xx.xxCustom Summary Functions
You can specify which summary functions to use, including custom ones.
Built-in Summary Functions
GeoTessera provides several built-in summary functions:
| Function | Description |
|---|---|
summary_mean |
Mean across all pixels |
summary_median |
Median across all pixels |
summary_sd |
Standard deviation per dimension |
summary_centroid |
Sample at geographic centroid |
summary_pixel_count |
Count of valid/total pixels |
summary_coverage |
Coverage statistics |
summary_quantile(probs) |
Factory for quantile summaries |
summary_random_sample(n, seed) |
Factory for random sampling |
# Use specific summary functions
result <- gt$summarize_region(
region = aba_north,
year = 2024,
summary_fns = list(
mean = summary_mean,
median = summary_median,
sd = summary_sd,
coverage = summary_coverage
)
)
# Standard deviation tells us about embedding variability
sd_embedding <- result$summaries$sd
summary(sd_embedding)Quantile Summaries
Use summary_quantile() to create a function that
computes specific quantiles:
# Create a function for 10th and 90th percentiles
result <- gt$summarize_region(
region = aba_north,
year = 2024,
summary_fns = list(
q10_90 = summary_quantile(c(0.10, 0.90))
)
)
# Returns 256 values (128 dimensions × 2 quantiles)
length(result$summaries$q10_90)Random Sampling
Use summary_random_sample() to extract random pixel
embeddings:
# Sample 50 random pixels (reproducible with seed)
result <- gt$summarize_region(
region = aba_north,
year = 2024,
summary_fns = list(
samples = summary_random_sample(n = 50, seed = 42)
)
)
# Returns a matrix: 50 samples × 128 dimensions
dim(result$summaries$samples)
# [1] 50 128Custom Summary Functions
Create your own summary function. It should accept:
-
embeddings: List of 3D arrays (height × width × 128) -
region: The input region (sf object or bbox) -
tiles_df: Tile metadata data frame
# Custom function: compute the embedding at the pixel with maximum brightness
# (using sum across channels as a proxy for "brightness")
summary_max_brightness <- function(embeddings, region = NULL, tiles_df = NULL) {
max_sum <- -Inf
max_embedding <- NULL
for (emb in embeddings) {
# Sum across channels for each pixel
pixel_sums <- apply(emb, c(1, 2), sum, na.rm = TRUE)
# Find max
max_idx <- which.max(pixel_sums)
if (pixel_sums[max_idx] > max_sum) {
max_sum <- pixel_sums[max_idx]
# Convert linear index to array indices
idx <- arrayInd(max_idx, dim(pixel_sums))
max_embedding <- emb[idx[1], idx[2], ]
}
}
names(max_embedding) <- paste0("dim_", seq_along(max_embedding))
max_embedding
}
# Use the custom function
result <- gt$summarize_region(
region = aba_north,
year = 2024,
summary_fns = list(
mean = summary_mean,
max_brightness = summary_max_brightness
)
)Processing Multiple Regions
When processing multiple regions, adjacent areas often share tiles.
GeoTessera provides an optimized
summarize_regions_streaming() function that:
- Builds a region-to-tile mapping - identifies which tiles each region needs
- Detects shared tiles - finds tiles used by multiple regions
- Optimizes download order - processes most-shared tiles first
- Downloads each tile only once - even if multiple regions need it
- Uses streaming statistics - constant memory regardless of region size
Efficient Multi-Region Processing
# Select adjacent LGAs that share tiles
selected_lgas <- c("Aba North", "Aba South", "Osisioma Ngwa", "Ugwunagbo")
lgas_subset <- nigeria_lgas[nigeria_lgas$adminName %in% selected_lgas, ]
# Process all regions efficiently
result <- summarize_regions_streaming(
gt = gt,
regions = lgas_subset,
year = 2024,
region_ids = lgas_subset$adminName,
sample_rate = 0.1, # Sample 10% of pixels for speed
mask_to_region = TRUE # Only include pixels inside each polygon
)
# View optimization statistics
result$metadata
# $n_regions
# [1] 4
#
# $n_unique_tiles
# [1] 11
#
# $tiles_downloaded
# [1] 11
#
# (4 tiles were shared between regions - downloaded only once!)The function returns:
-
summaries: Named list of mean embeddings (128-dim vector per region) -
pixel_counts: Number of pixels processed per region -
metadata: Processing statistics
# Access results for a specific region
aba_north_embedding <- result$summaries[["Aba North"]]
length(aba_north_embedding)
# [1] 128
# View pixel counts
result$pixel_counts
# Aba North Aba South Osisioma Ngwa Ugwunagbo
# 18800 43538 175735 126769
# Combine into a matrix for analysis
embedding_matrix <- do.call(rbind, result$summaries)
dim(embedding_matrix)
# [1] 4 128Comparison: Sequential vs Optimized
For adjacent regions, the optimized approach is significantly faster:
Sequential processing (4 LGAs separately):
- Each LGA downloads its own tiles
- Shared tiles downloaded multiple times
- Total: ~20+ tile downloads
Optimized streaming (summarize_regions_streaming):
- Builds tile sharing graph
- Downloads each unique tile once
- Total: 11 tile downloads (4 shared)Large-Scale Processing
For processing many regions (e.g., all LGAs in a country), use sampling to reduce processing time:
# Process all LGAs in a state
abia_lgas <- nigeria_lgas[grepl("^NG001", nigeria_lgas$adminCode), ]
result <- summarize_regions_streaming(
gt = gt,
regions = abia_lgas,
year = 2024,
region_ids = abia_lgas$adminName,
sample_rate = 0.05, # 5% sampling for very large jobs
mask_to_region = TRUE
)Creating a Summary Data Frame
Combine embeddings with administrative metadata:
# Create data frame with LGA info and embeddings
embedding_matrix <- do.call(rbind, result$summaries)
lga_embeddings <- data.frame(
adminName = names(result$summaries),
embedding_matrix
)
# Add area information
lga_embeddings$area_km2 <- as.numeric(st_area(lgas_subset)) / 1e6
# Now you can use this for clustering, classification, etc.
head(lga_embeddings[, 1:5])Controlling Masking Behavior
By default, pixels outside the region boundary are masked (set to NA). You can disable this to include all pixels within the bounding box:
# Without masking - uses rectangular bounding box
result_unmasked <- gt$summarize_region(
region = aba_north,
year = 2024,
mask_to_region = FALSE
)
# With masking (default) - only pixels inside the polygon
result_masked <- gt$summarize_region(
region = aba_north,
year = 2024,
mask_to_region = TRUE
)
# Compare coverage
result_unmasked$summaries$coverage["n_valid_pixels"]
result_masked$summaries$coverage["n_valid_pixels"]
# Masked version will have fewer valid pixelsUsing Bounding Boxes
You can also use simple bounding boxes instead of sf objects:
# Define a bounding box (xmin, ymin, xmax, ymax)
bbox <- c(7.33, 5.08, 7.40, 5.14)
result <- gt$summarize_region(
region = bbox,
year = 2024
)
# Note: masking is not applied for bounding boxes
result$metadata$is_masked
# [1] FALSEPerformance Considerations
Temporary File Management
Downloaded tiles are stored in a temporary directory and
automatically cleaned up when summarize_region() completes.
If you need to keep the tiles:
Large Regions (Memory-Efficient Streaming)
For very large regions (many tiles), the standard
summarize_region() may run out of memory because it loads
all embeddings before computing statistics.
Use summarize_region_streaming() for memory-efficient
processing of large single regions:
# Abaji LGA requires 36 tiles - too large for standard approach
abaji <- nigeria_lgas[nigeria_lgas$adminName == "Abaji", ]
# Use streaming approach - processes one tile at a time
result <- gt$summarize_region_streaming(
region = abaji,
year = 2024,
sample_rate = 0.1, # Sample 10% of pixels
mask_to_region = FALSE # Skip masking for speed
)
# Results
result$mean # 128-dimensional mean embedding
result$n_pixels # Number of pixels processed
result$metadata$processing_time_secsThe streaming approach:
- Constant memory: Processes one tile at a time using Welford’s online algorithm
-
Sampling support: Reduce processing time with
sample_rate < 1.0 - No memory issues: Successfully handles regions with 30+ tiles
Alternative: Point Sampling Within Polygon
Another memory-efficient approach is to sample random points within the polygon and fetch their embeddings:
# Sample 1000 random points inside the polygon
set.seed(42)
sample_points <- st_sample(abaji, size = 1000)
coords <- st_coordinates(sample_points)
points_df <- data.frame(lon = coords[, 1], lat = coords[, 2])
# Fetch embeddings at those points
embeddings_df <- gt$sample_embeddings_at_points(points_df, year = 2024)
# Compute mean (excluding lon/lat columns)
mean_embedding <- colMeans(embeddings_df[, -(1:2)], na.rm = TRUE)Example: Clustering LGAs by Land Cover Similarity
Here’s a complete example that clusters Nigerian LGAs based on embedding similarity, using the optimized multi-region streaming approach:
library(GeoTessera)
library(sf)
# Load shapefile
nigeria_lgas <- st_read("shapefiles/nigeria_admin2_shapefile.shp", quiet = TRUE)
# Calculate areas and select small LGAs for demo
nigeria_lgas$area_km2 <- as.numeric(st_area(nigeria_lgas)) / 1e6
small_lgas <- nigeria_lgas[nigeria_lgas$area_km2 < 100, ]
small_lgas <- small_lgas[1:20, ] # Take first 20 for demo
# Initialize client
gt <- geotessera()
# Compute mean embeddings for all LGAs efficiently
# This optimizes tile downloads when regions share tiles
result <- summarize_regions_streaming(
gt = gt,
regions = small_lgas,
year = 2024,
region_ids = small_lgas$adminName,
sample_rate = 0.1,
mask_to_region = TRUE
)
# Create embedding matrix from results
emb_matrix <- do.call(rbind, result$summaries)
# Hierarchical clustering
dist_matrix <- dist(emb_matrix)
hc <- hclust(dist_matrix, method = "ward.D2")
# Plot dendrogram
plot(hc, main = "LGA Clustering by Embedding Similarity",
xlab = "", sub = "")
# Cut into clusters
clusters <- cutree(hc, k = 4)
# Add cluster assignments back to spatial data
small_lgas$cluster <- clusters[small_lgas$adminName]
# Plot clusters on map
plot(small_lgas["cluster"], main = "LGA Clusters")Summary
The summarize_region() method provides a convenient
pipeline for:
- Downloading embeddings for arbitrary regions
- Masking to irregular boundaries from shapefiles
- Computing flexible summary statistics
- Cleaning up temporary files automatically
This enables analysis workflows where you need region-level embeddings rather than full pixel-level data, such as:
- Administrative unit characterization
- Regional clustering and classification
- Time series analysis at the regional level
- Feature extraction for machine learning models