Machine Learning with Tessera Embeddings
Source:vignettes/machine-learning.Rmd
machine-learning.RmdIntroduction
Tessera embeddings are designed to be used as features for machine learning tasks. The 128-dimensional embedding vectors capture rich semantic information about land cover, vegetation, urban areas, and other environmental features derived from Sentinel-1 and Sentinel-2 satellite imagery.
This vignette demonstrates common machine learning workflows using the Cambridge region (4 tiles, ~400 MB) as a practical example.
Demonstration with Simulated Data
Before working with real data, let’s demonstrate the machine learning workflows using simulated embeddings. This helps understand the structure without requiring downloads.
Simulated 128-Channel Embeddings
set.seed(42)
n_samples <- 500
# Simulate 128-dimensional embeddings with 3 land cover types
# Each class has distinct patterns in embedding space
generate_class_embeddings <- function(n, class_id) {
# Base pattern varies by class
base <- switch(class_id,
"urban" = rnorm(128, mean = 0.5, sd = 0.1),
"vegetation" = rnorm(128, mean = -0.3, sd = 0.15),
"water" = rnorm(128, mean = 0.1, sd = 0.05)
)
# Generate samples with class-specific variation
samples <- matrix(nrow = n, ncol = 128)
for (i in 1:n) {
samples[i, ] <- base + rnorm(128, sd = 0.2)
}
samples
}
# Generate data for each class
urban_emb <- generate_class_embeddings(200, "urban")
veg_emb <- generate_class_embeddings(200, "vegetation")
water_emb <- generate_class_embeddings(100, "water")
# Combine into training data
X <- rbind(urban_emb, veg_emb, water_emb)
y <- factor(c(rep("urban", 200), rep("vegetation", 200), rep("water", 100)))
cat("Training data dimensions:", dim(X), "\n")
#> Training data dimensions: 500 128
cat("Class distribution:\n")
#> Class distribution:
print(table(y))
#> y
#> urban vegetation water
#> 200 200 100PCA Visualization of Embeddings
# Apply PCA to visualize the 128-dimensional embeddings
pca_result <- prcomp(X, center = TRUE, scale. = TRUE)
# Variance explained
var_explained <- pca_result$sdev^2 / sum(pca_result$sdev^2)
# Create visualization data
pca_df <- data.frame(
PC1 = pca_result$x[, 1],
PC2 = pca_result$x[, 2],
PC3 = pca_result$x[, 3],
Class = y
)
# Plot PC1 vs PC2
ggplot(pca_df, aes(x = PC1, y = PC2, color = Class)) +
geom_point(alpha = 0.6, size = 2) +
scale_color_manual(values = c("urban" = "#E41A1C",
"vegetation" = "#4DAF4A",
"water" = "#377EB8")) +
labs(title = "PCA of Simulated Tessera Embeddings",
subtitle = sprintf("PC1: %.1f%% var, PC2: %.1f%% var",
var_explained[1]*100, var_explained[2]*100),
x = "Principal Component 1",
y = "Principal Component 2") +
theme_minimal() +
theme(legend.position = "bottom")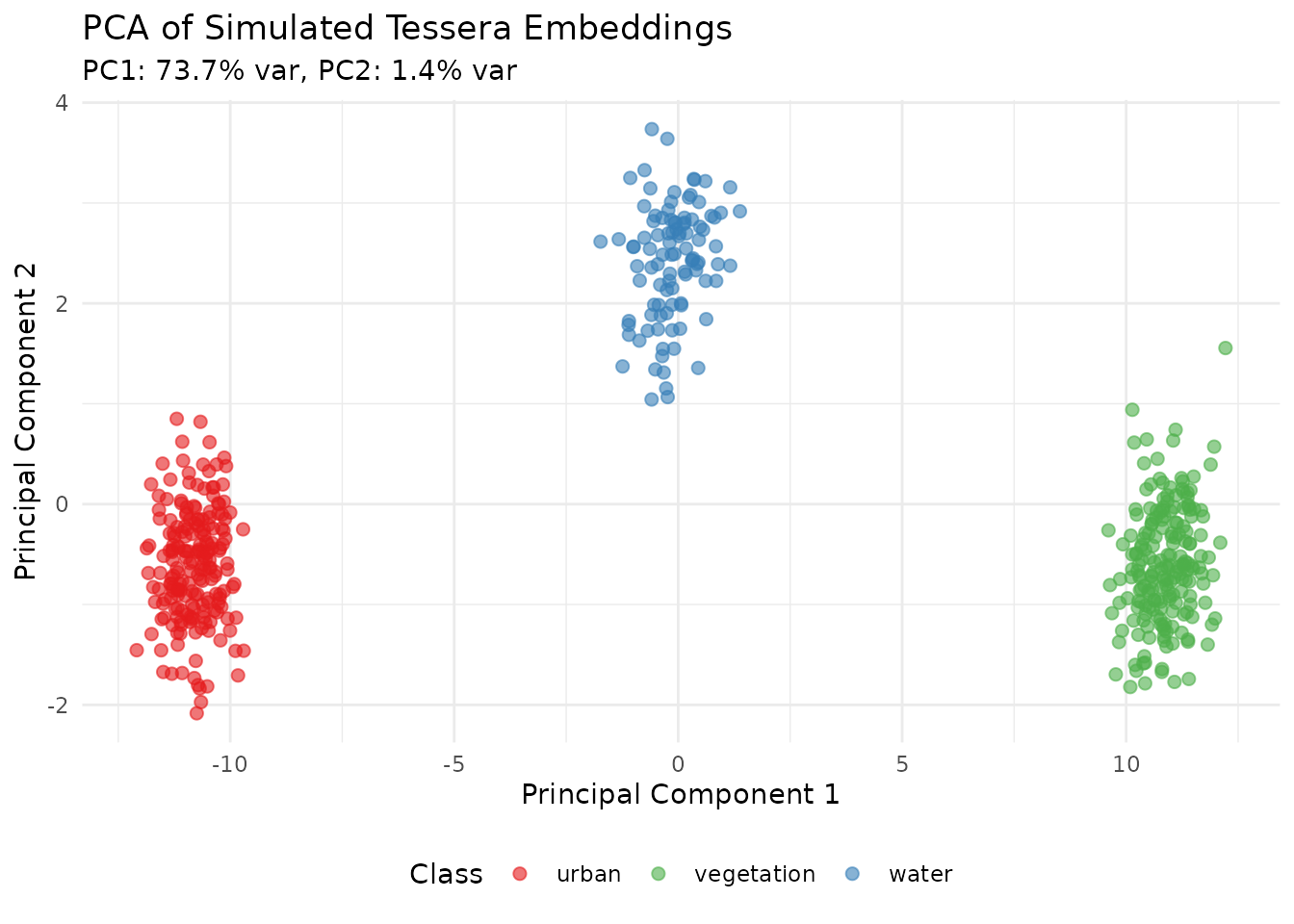
Scree Plot
scree_df <- data.frame(
PC = 1:20,
Variance = var_explained[1:20],
Cumulative = cumsum(var_explained[1:20])
)
ggplot(scree_df, aes(x = PC)) +
geom_bar(aes(y = Variance), stat = "identity", fill = "steelblue", alpha = 0.7) +
geom_line(aes(y = Cumulative), color = "red", linewidth = 1) +
geom_point(aes(y = Cumulative), color = "red", size = 2) +
geom_hline(yintercept = 0.95, linetype = "dashed", color = "gray50") +
scale_y_continuous(
name = "Variance Explained",
sec.axis = sec_axis(~., name = "Cumulative Variance")
) +
labs(title = "PCA Scree Plot",
subtitle = "Dashed line shows 95% cumulative variance threshold",
x = "Principal Component") +
theme_minimal()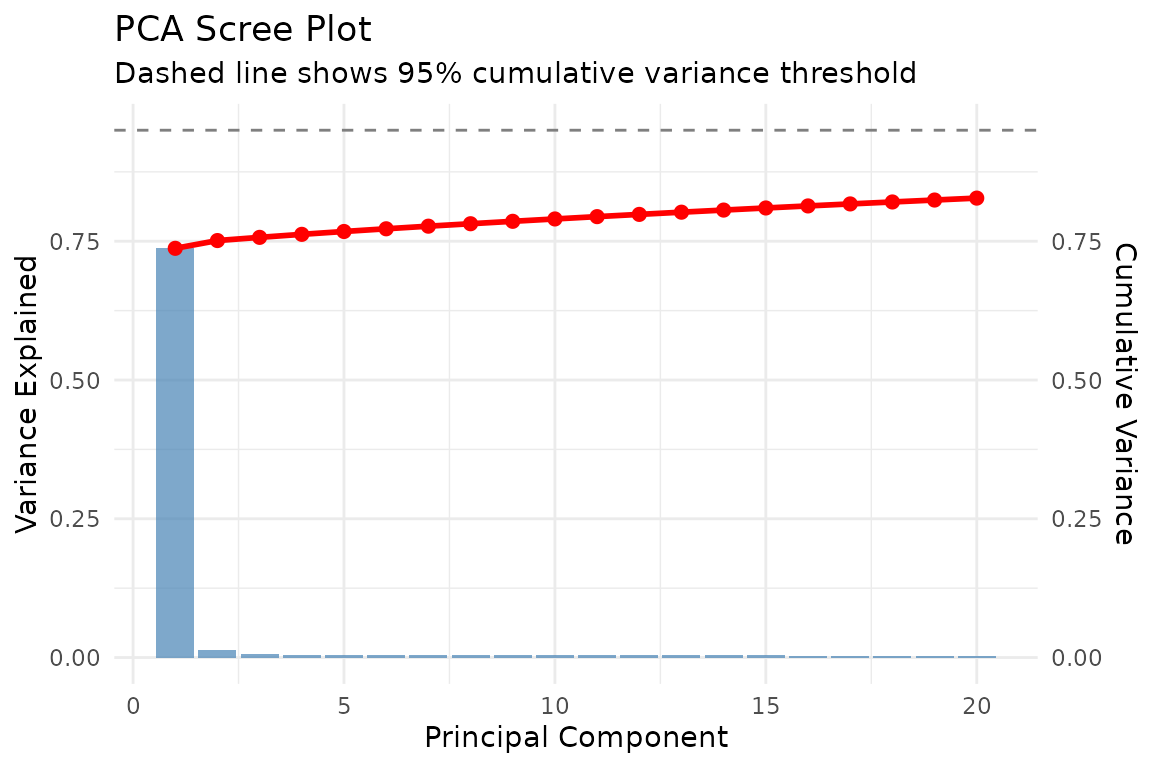
K-Means Clustering
# K-means clustering (unsupervised)
set.seed(123)
kmeans_result <- kmeans(X, centers = 3, nstart = 25)
pca_df$Cluster <- factor(kmeans_result$cluster)
# Compare clusters to actual classes
ggplot(pca_df, aes(x = PC1, y = PC2)) +
geom_point(aes(color = Cluster, shape = Class), alpha = 0.6, size = 2) +
scale_color_brewer(palette = "Set1") +
labs(title = "K-Means Clustering vs True Classes",
subtitle = "Colors = K-means clusters, Shapes = True classes",
x = "PC1", y = "PC2") +
theme_minimal() +
theme(legend.position = "bottom")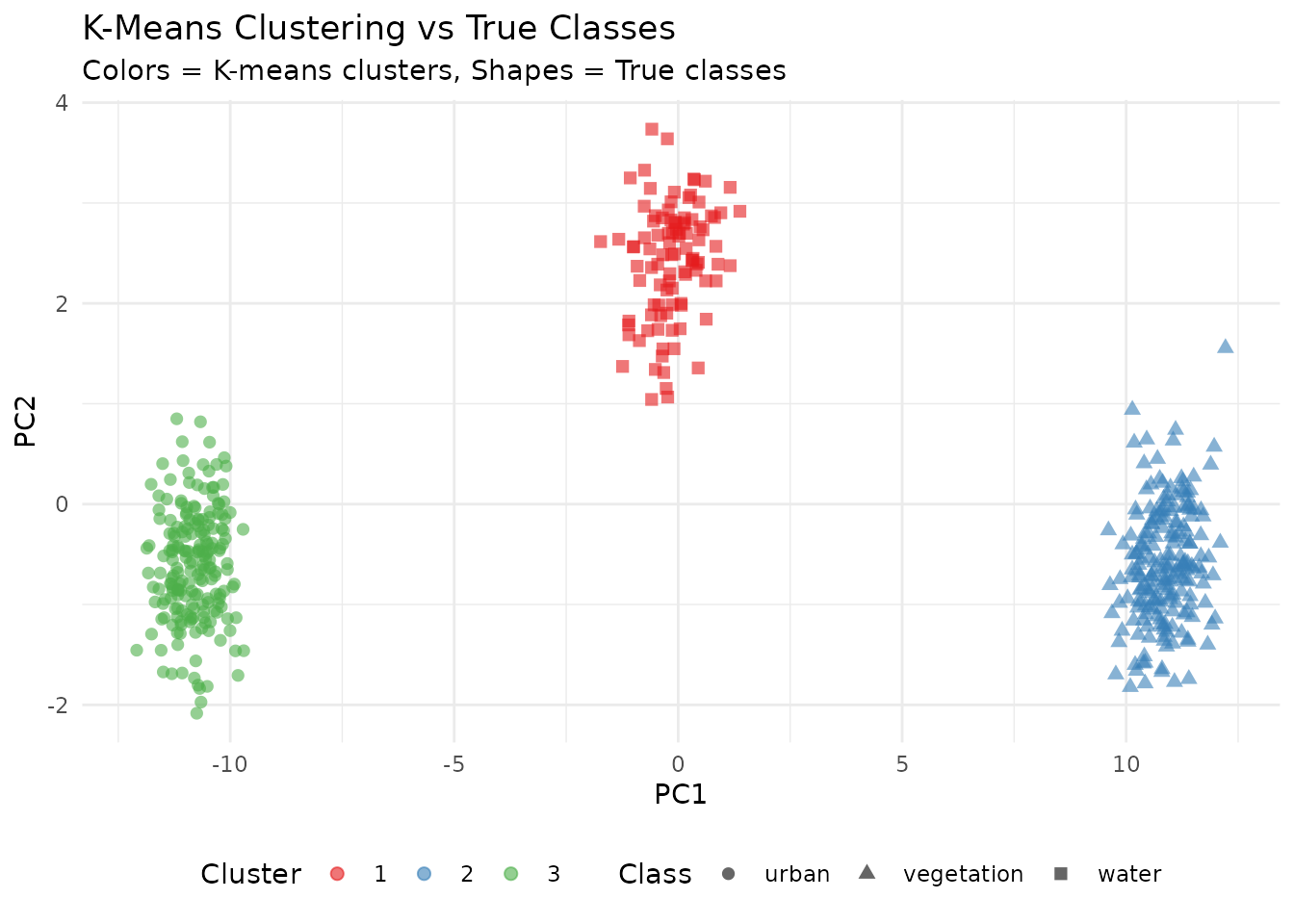
Classification Confusion Matrix
# Simple classification assessment using cluster assignment
# Map clusters to most common class
cluster_class_map <- sapply(1:3, function(k) {
names(which.max(table(y[kmeans_result$cluster == k])))
})
predicted <- factor(cluster_class_map[kmeans_result$cluster], levels = levels(y))
conf_matrix <- table(Predicted = predicted, Actual = y)
# Convert to data frame for plotting
conf_df <- as.data.frame(conf_matrix)
ggplot(conf_df, aes(x = Actual, y = Predicted, fill = Freq)) +
geom_tile() +
geom_text(aes(label = Freq), color = "white", size = 5) +
scale_fill_gradient(low = "steelblue", high = "darkred") +
labs(title = "Classification Confusion Matrix",
subtitle = "Cluster-based class assignment") +
theme_minimal() +
theme(legend.position = "none")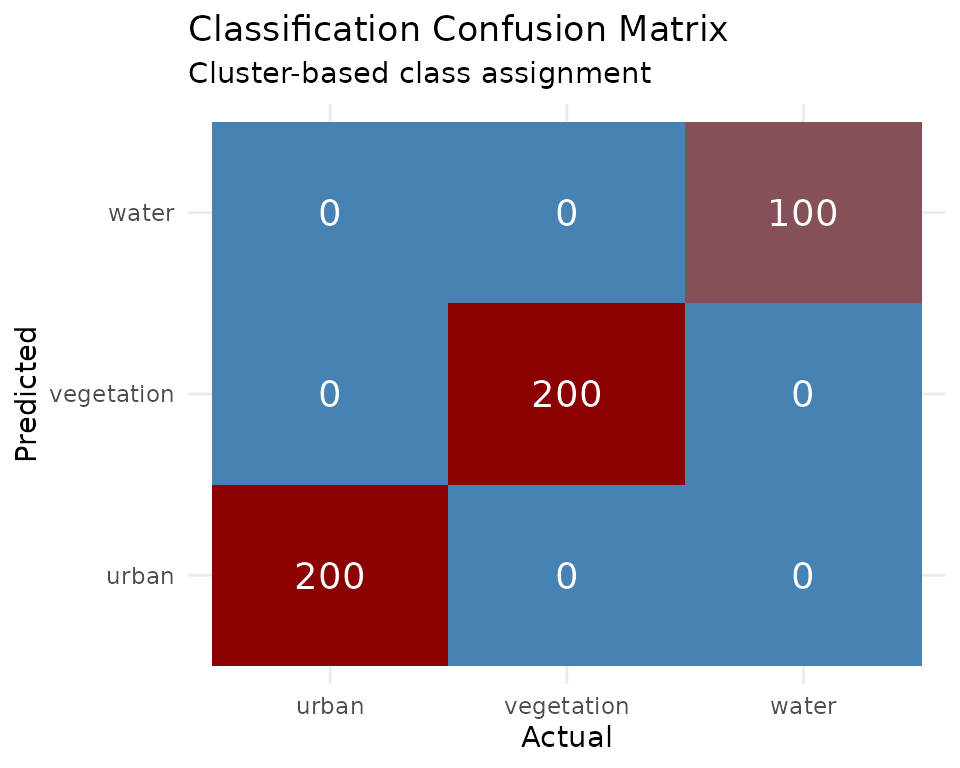
Feature Importance (Simulated)
# Simulate feature importance (as would come from Random Forest)
set.seed(456)
importance_df <- data.frame(
Band = paste0("Band_", 1:128),
Importance = abs(rnorm(128, mean = 0.02, sd = 0.015))
)
importance_df <- importance_df[order(-importance_df$Importance), ]
importance_df$Rank <- 1:128
# Plot top 20
ggplot(importance_df[1:20, ], aes(x = reorder(Band, Importance), y = Importance)) +
geom_bar(stat = "identity", fill = "steelblue") +
coord_flip() +
labs(title = "Top 20 Most Important Embedding Bands",
subtitle = "Simulated Random Forest variable importance",
x = "Embedding Band", y = "Importance") +
theme_minimal()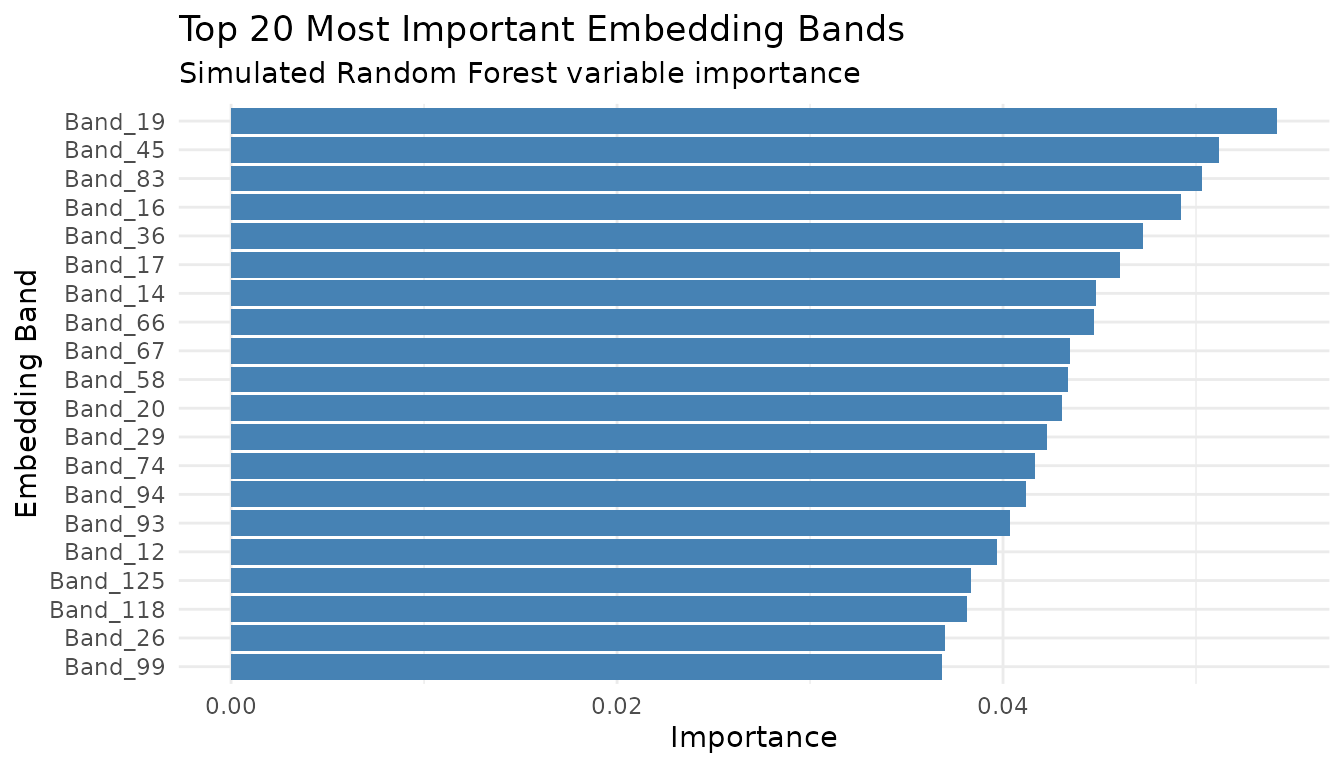
Download Cambridge Region Data
First, let’s download the Cambridge region embeddings which we’ll use throughout this vignette.
# Create client
gt <- geotessera()
# Cambridge region bounding box (4 tiles)
cambridge_bbox <- list(
xmin = 0.086174,
ymin = 52.183432,
xmax = 0.151062,
ymax = 52.206318
)
# Get tile metadata
tiles <- gt$get_tiles(bbox = cambridge_bbox, year = 2024)
print(paste("Tiles:", nrow(tiles)))
# Expected: 4 tiles
# Export as GeoTIFFs
output_dir <- "cambridge_tiles"
paths <- gt$export_embedding_geotiffs(
tiles = tiles,
output_dir = output_dir,
compress = "lzw",
progress = TRUE
)
# Discover and load tiles
tile_list <- discover_tiles(output_dir, format = "geotiff")
print(paste("Loaded tiles:", length(tile_list)))Land Cover Classification
A common use case is classifying land cover types using labeled training data.
Preparing Training Data with Real Embeddings
# Sample training locations across Cambridge
# These represent different land cover types in the region
training_sites <- data.frame(
lon = c(
0.10, 0.11, 0.095, 0.12, 0.105, # Urban/built-up
0.09, 0.14, 0.135, 0.088, 0.13, # Green spaces/parks
0.115, 0.125 # Water/wetland
),
lat = c(
52.19, 52.195, 52.188, 52.20, 52.192, # Urban
52.185, 52.20, 52.195, 52.19, 52.185, # Green
52.205, 52.198 # Water
),
land_cover = c(
rep("urban", 5),
rep("green", 5),
rep("water", 2)
)
)
# Sample embeddings at training locations
training_embeddings <- gt$sample_embeddings_at_points(
points = training_sites,
year = 2024,
download = TRUE
)
# Combine with labels
X_train <- as.matrix(training_embeddings[, 3:130]) # embedding_1 to embedding_128
y_train <- as.factor(training_sites$land_cover)
print(dim(X_train))
# [1] 12 128
print(table(y_train))
# green urban water
# 5 5 2Training a Random Forest Classifier
library(randomForest)
# Train random forest
set.seed(42)
rf_model <- randomForest(
x = X_train,
y = y_train,
ntree = 500,
importance = TRUE
)
print(rf_model)
# Variable importance - which embedding bands are most predictive?
importance_df <- as.data.frame(importance(rf_model))
importance_df$band <- rownames(importance_df)
importance_df <- importance_df[order(-importance_df$MeanDecreaseAccuracy), ]
print(head(importance_df, 20))Predicting on New Locations
# Sample embeddings at new locations within Cambridge
new_sites <- data.frame(
lon = c(0.09, 0.11, 0.13, 0.10),
lat = c(52.19, 52.20, 52.185, 52.195)
)
new_embeddings <- gt$sample_embeddings_at_points(
points = new_sites,
year = 2024
)
X_new <- as.matrix(new_embeddings[, 3:130])
# Make predictions
predictions <- predict(rf_model, X_new)
probabilities <- predict(rf_model, X_new, type = "prob")
results <- cbind(new_sites, prediction = predictions, probabilities)
print(results)Classifying an Entire Tile
# Load a Cambridge tile
tile <- gt$download_tile(lon = 0.1, lat = 52.2, year = 2024)
# Load embedding as array
embedding <- tile$load_embedding()
dims <- dim(embedding) # (1111, 1111, 128)
# Reshape to 2D matrix for prediction
embedding_2d <- matrix(embedding, nrow = dims[1] * dims[2], ncol = dims[3])
# Predict (1.2M pixels)
pixel_predictions <- predict(rf_model, embedding_2d)
# Reshape back to image
classified <- matrix(
as.integer(pixel_predictions),
nrow = dims[1],
ncol = dims[2]
)
# Create raster
bounds <- tile$get_bounds()
r_classified <- rast(
nrows = dims[1],
ncols = dims[2],
xmin = bounds$xmin,
xmax = bounds$xmax,
ymin = bounds$ymin,
ymax = bounds$ymax,
crs = "EPSG:4326"
)
values(r_classified) <- as.vector(t(classified))
# Save result
writeRaster(r_classified, "cambridge_land_cover.tif", overwrite = TRUE)
# Plot
plot(r_classified, main = "Land Cover Classification - Cambridge")Clustering Analysis
Use unsupervised learning to discover natural groupings in Cambridge embeddings.
K-Means Clustering
# Sample random points across Cambridge region
set.seed(123)
n_samples <- 2000
random_points <- data.frame(
lon = runif(n_samples, cambridge_bbox$xmin, cambridge_bbox$xmax),
lat = runif(n_samples, cambridge_bbox$ymin, cambridge_bbox$ymax)
)
# Get embeddings
embeddings <- gt$sample_embeddings_at_points(random_points, year = 2024)
X <- as.matrix(embeddings[, 3:130])
# Remove any NA rows
valid_rows <- complete.cases(X)
X_valid <- X[valid_rows, ]
points_valid <- random_points[valid_rows, ]
print(paste("Valid samples:", nrow(X_valid)))
# K-means clustering
set.seed(42)
k <- 5
kmeans_result <- kmeans(X_valid, centers = k, nstart = 25, iter.max = 100)
# Add cluster assignments
points_valid$cluster <- as.factor(kmeans_result$cluster)
# Visualize
library(ggplot2)
ggplot(points_valid, aes(x = lon, y = lat, color = cluster)) +
geom_point(size = 1, alpha = 0.7) +
coord_fixed() +
theme_minimal() +
labs(
title = "K-Means Clustering of Cambridge Embeddings",
subtitle = paste("k =", k, "clusters,", nrow(points_valid), "samples")
)Hierarchical Clustering
# Use subset for hierarchical clustering (memory intensive)
set.seed(42)
subset_idx <- sample(nrow(X_valid), min(500, nrow(X_valid)))
X_subset <- X_valid[subset_idx, ]
# Compute distance matrix
dist_matrix <- dist(X_subset)
# Hierarchical clustering
hc <- hclust(dist_matrix, method = "ward.D2")
# Plot dendrogram
plot(hc, labels = FALSE, main = "Hierarchical Clustering - Cambridge Embeddings")
# Cut tree at desired number of clusters
clusters <- cutree(hc, k = 5)Dimensionality Reduction
PCA for Visualization
# Apply PCA to Cambridge embeddings
pca_result <- prcomp(X_valid, center = TRUE, scale. = TRUE)
# Variance explained
var_explained <- pca_result$sdev^2 / sum(pca_result$sdev^2)
cumvar <- cumsum(var_explained)
# How many components for 95% variance?
n_95 <- which(cumvar >= 0.95)[1]
print(paste("Components for 95% variance:", n_95))
# Plot variance explained
plot(var_explained[1:50], type = "b",
xlab = "Principal Component", ylab = "Variance Explained",
main = "PCA Scree Plot - Cambridge Embeddings")
# Visualize first two PCs with cluster coloring
pca_df <- data.frame(
PC1 = pca_result$x[, 1],
PC2 = pca_result$x[, 2],
cluster = points_valid$cluster
)
ggplot(pca_df, aes(x = PC1, y = PC2, color = cluster)) +
geom_point(alpha = 0.5, size = 0.5) +
theme_minimal() +
labs(title = "PCA of Cambridge Embeddings")UMAP for Nonlinear Visualization
library(uwot)
# UMAP embedding
set.seed(42)
umap_result <- umap(
X_valid,
n_neighbors = 15,
min_dist = 0.1,
n_components = 2,
metric = "euclidean"
)
# Visualize
umap_df <- data.frame(
UMAP1 = umap_result[, 1],
UMAP2 = umap_result[, 2],
cluster = points_valid$cluster
)
ggplot(umap_df, aes(x = UMAP1, y = UMAP2, color = cluster)) +
geom_point(alpha = 0.5, size = 0.5) +
theme_minimal() +
labs(title = "UMAP Projection of Cambridge Embeddings")Change Detection
Compare embeddings from different years to detect changes in Cambridge.
Temporal Difference Analysis
# Sample locations for change analysis
analysis_points <- data.frame(
lon = seq(cambridge_bbox$xmin, cambridge_bbox$xmax, length.out = 100),
lat = rep(52.195, 100)
)
# Get embeddings for two different years
embeddings_2020 <- gt$sample_embeddings_at_points(analysis_points, year = 2020)
embeddings_2024 <- gt$sample_embeddings_at_points(analysis_points, year = 2024)
# Calculate change magnitude (Euclidean distance in embedding space)
X_2020 <- as.matrix(embeddings_2020[, 3:130])
X_2024 <- as.matrix(embeddings_2024[, 3:130])
change_magnitude <- sqrt(rowSums((X_2024 - X_2020)^2))
# Create results
change_df <- data.frame(
lon = analysis_points$lon,
lat = analysis_points$lat,
change = change_magnitude
)
# Visualize
ggplot(change_df, aes(x = lon, y = change)) +
geom_line() +
geom_point(aes(color = change)) +
scale_color_viridis_c() +
theme_minimal() +
labs(
title = "Embedding Change Magnitude (2020-2024)",
subtitle = "Cambridge transect",
x = "Longitude",
y = "Change Magnitude (Euclidean distance)"
)Change Maps
# Download tiles for two years
tile_2020 <- gt$download_tile(lon = 0.1, lat = 52.2, year = 2020)
tile_2024 <- gt$download_tile(lon = 0.1, lat = 52.2, year = 2024)
# Load embeddings
emb_2020 <- tile_2020$load_embedding()
emb_2024 <- tile_2024$load_embedding()
# Calculate per-pixel change magnitude
change_array <- sqrt(rowSums((emb_2024 - emb_2020)^2, dims = 2))
# Create raster
bounds <- tile_2020$get_bounds()
r_change <- rast(
nrows = dim(change_array)[1],
ncols = dim(change_array)[2],
xmin = bounds$xmin,
xmax = bounds$xmax,
ymin = bounds$ymin,
ymax = bounds$ymax,
crs = "EPSG:4326"
)
values(r_change) <- as.vector(t(change_array))
# Plot
plot(r_change, main = "Change Detection 2020-2024 - Cambridge",
col = hcl.colors(50, "YlOrRd", rev = TRUE))Regression Tasks
Predict continuous variables using Cambridge embeddings.
Gradient Boosting Regression
library(xgboost)
# Example: Predict a continuous variable from embeddings
# (Using simulated target for demonstration - replace with real data)
training_with_target <- data.frame(
lon = c(0.10, 0.095, 0.11, 0.12, 0.085, 0.13, 0.09, 0.115),
lat = c(52.19, 52.185, 52.20, 52.195, 52.188, 52.19, 52.20, 52.192),
target_value = c(120, 85, 150, 140, 90, 110, 95, 135)
)
# Get embeddings
train_emb <- gt$sample_embeddings_at_points(training_with_target, year = 2024)
X <- as.matrix(train_emb[, 3:130])
y <- training_with_target$target_value
# Train XGBoost
dtrain <- xgb.DMatrix(data = X, label = y)
params <- list(
objective = "reg:squarederror",
eta = 0.1,
max_depth = 4
)
xgb_model <- xgb.train(
params = params,
data = dtrain,
nrounds = 50,
verbose = 0
)
# Feature importance
importance <- xgb.importance(model = xgb_model)
xgb.plot.importance(importance[1:20, ], main = "Top 20 Important Embedding Bands")
# Predict on new data
new_points <- data.frame(lon = c(0.1, 0.12), lat = c(52.19, 52.20))
new_embeddings <- gt$sample_embeddings_at_points(new_points, year = 2024)
X_new <- as.matrix(new_embeddings[, 3:130])
predictions <- predict(xgb_model, X_new)
print(paste("Predictions:", paste(round(predictions, 1), collapse = ", ")))Deep Learning with torch
Use Cambridge embeddings with deep learning frameworks.
Preparing Data for torch
library(torch)
# Convert embeddings to torch tensors
X_tensor <- torch_tensor(X_valid, dtype = torch_float())
y_tensor <- torch_tensor(as.integer(points_valid$cluster), dtype = torch_long())
# Create dataset
dataset <- tensor_dataset(X_tensor, y_tensor)
# Create dataloader
dataloader <- dataloader(dataset, batch_size = 32, shuffle = TRUE)Simple Neural Network Classifier
# Define model
net <- nn_module(
"EmbeddingClassifier",
initialize = function(input_size = 128, hidden_size = 64, num_classes = 5) {
self$fc1 <- nn_linear(input_size, hidden_size)
self$relu <- nn_relu()
self$dropout <- nn_dropout(p = 0.3)
self$fc2 <- nn_linear(hidden_size, hidden_size %/% 2)
self$fc3 <- nn_linear(hidden_size %/% 2, num_classes)
},
forward = function(x) {
x %>%
self$fc1() %>%
self$relu() %>%
self$dropout() %>%
self$fc2() %>%
self$relu() %>%
self$dropout() %>%
self$fc3()
}
)
# Instantiate model
model <- net(input_size = 128, hidden_size = 64, num_classes = k)
# Training setup
optimizer <- optim_adam(model$parameters, lr = 0.001)
loss_fn <- nn_cross_entropy_loss()
# Training loop
epochs <- 10
for (epoch in 1:epochs) {
model$train()
total_loss <- 0
n_batches <- 0
coro::loop(for (batch in dataloader) {
optimizer$zero_grad()
output <- model(batch[[1]])
loss <- loss_fn(output, batch[[2]])
loss$backward()
optimizer$step()
total_loss <- total_loss + loss$item()
n_batches <- n_batches + 1
})
cat(sprintf("Epoch %d, Avg Loss: %.4f\n", epoch, total_loss / n_batches))
}Working with All Cambridge Tiles
Process all 4 Cambridge tiles systematically.
Batch Processing
# Process each tile
results <- list()
for (i in seq_len(nrow(tiles))) {
tile_info <- tiles[i, ]
# Download tile
tile <- gt$download_tile(
lon = tile_info$lon,
lat = tile_info$lat,
year = tile_info$year,
progress = FALSE
)
# Load embedding
embedding <- tile$load_embedding()
# Calculate summary statistics per tile
results[[i]] <- data.frame(
grid_name = tile$get_grid_name(),
lon = tile_info$lon,
lat = tile_info$lat,
mean_emb = mean(embedding, na.rm = TRUE),
sd_emb = sd(embedding, na.rm = TRUE),
min_emb = min(embedding, na.rm = TRUE),
max_emb = max(embedding, na.rm = TRUE)
)
cat(sprintf("Processed tile %d/%d: %s\n", i, nrow(tiles), tile$get_grid_name()))
}
# Combine results
summary_df <- do.call(rbind, results)
print(summary_df)Parallel Processing
library(parallel)
library(foreach)
library(doParallel)
# Set up parallel backend
n_cores <- min(detectCores() - 1, nrow(tiles))
cl <- makeCluster(n_cores)
registerDoParallel(cl)
# Process tiles in parallel
results <- foreach(
i = 1:nrow(tiles),
.packages = c("GeoTessera", "terra"),
.combine = rbind
) %dopar% {
gt_local <- geotessera()
tile_info <- tiles[i, ]
tile <- gt_local$download_tile(
lon = tile_info$lon,
lat = tile_info$lat,
year = tile_info$year,
progress = FALSE
)
embedding <- tile$load_embedding()
data.frame(
grid_name = tile$get_grid_name(),
mean_emb = mean(embedding, na.rm = TRUE),
sd_emb = sd(embedding, na.rm = TRUE)
)
}
stopCluster(cl)
print(results)Model Persistence
Save and load trained models for reuse.
# Save random forest model
saveRDS(rf_model, "cambridge_rf_model.rds")
# Load model
rf_model_loaded <- readRDS("cambridge_rf_model.rds")
# Save XGBoost model
xgb.save(xgb_model, "cambridge_xgb.model")
# Load XGBoost model
xgb_loaded <- xgb.load("cambridge_xgb.model")
# Save torch model
torch_save(model, "cambridge_classifier.pt")
# Load torch model
model_loaded <- torch_load("cambridge_classifier.pt")Best Practices for ML with Tessera Embeddings
Start with Cambridge: The 4-tile Cambridge region (~400 MB) is ideal for prototyping and testing ML workflows.
Normalize if needed: While embeddings are pre-scaled, additional normalization (z-score) may improve some models.
Spatial cross-validation: Nearby points are correlated. Use spatial cross-validation (e.g., leave-one-tile-out) for reliable evaluation.
Feature selection: Not all 128 bands may be relevant. Use importance scores to identify predictive bands.
Balance classes: For classification, ensure balanced class representation or use class weights.
Multi-year features: When available, combining embeddings from multiple years can capture temporal patterns.
Validate spatially: Test models on separate geographic regions to ensure generalization.